Der Workflowy Zeitstrahl
Ich benutze seit knapp zwei Jahren Workflowy als Notiz-App und seit einem knappen Jahr Frank Degenaars Zeitstrahlmethode (The Workflowy Timeline) als Notizsystem. Die meisten von uns schreiben in der einen oder anderen Form Notizen und früher oder später möchte man diese Notizen dann auch irgendwie organisieren – dann sind wir beim Notizsystem. Es gibt eine Fülle an solchen Systemen, ich kenne mich da nicht besonders aus, las nur David Allens Getting Things Done und hatte es dann in Teilen auch umgesetzt. Alle diese Systeme wollen natürlich nicht nur Notizen organisieren, sondern uns auch dabei helfen, Aufgaben und Projekte effizient und zur rechten Zeit zu erledigen, eben getting things done. Ich habe über die Jahre einiges durchprobiert und der Workflowy Zeitstrahl ist das System das für mich bisher und derzeit am besten funktioniert.
Workflowy ist eine der Millionen Notiz-Apps auf dem Planeten. Workflowy ist genau nur eine Liste an Stichpunkten, die unendlich tief verschachtelt sein kann. Ein Listeneintrag kann also eine Unterliste enthalten, die wiederum Unterlisten enthalten kann. Man kann in jeden Stichpunkt hineinzoomen, wodurch dieser zur Überschrift, die Unterliste des Stichpunkts zum eigentlichen Dokument wird. Unterlisten können zusammengefaltet werden, um die Übersicht zu wahren.

Das Bestechende an Workflowy ist also, dass es konzeptionell extrem einfach ist. Praktisch brauche ich nicht darüber nachdenken, wie ich meine Notizen in Dateien oder Dokumente sortiere, ich brauche diese auch nicht anlegen. Das ist eine große Sache. Zudem sehe ich immer nur das, was ich gerade sehen will, was für mich gerade relevant ist – durch Hineinzoomen und Zusammenfalten.
Natürlich bietet Workflowy eine Vielzahl an weiteren Funktionen, wie Tags, Personen (Mentions), Boards, gespiegelte Listeneinträge. Die meisten davon benutze ich nicht oder nur rudimentär. Aus meiner Sicht sollte Workflowy einfach bleiben. Was Workflowy nicht kann ist Prosa. Also geschriebener, fließender Text wie dieser Artikel. Einen solchen Artikel als Stichpunkte einer Liste abzubilden geht zwar, ist aber ästhetisch sehr unbefriedigend. Einerseits ist das okay, dafür ist Workflowy nicht gemacht. Andererseits wäre es fantastisch, wenn ich Artikel und Tagebücher angenehm in Workflowy schreiben könnte. Dann wäre wirklich alles, was ich schreibe, an nur einem Ort.
Workflowy hat im Übrigen auch gute mobile Apps mit gut funktionierender Synchronisation. Ich vermute, das haben fast alle Notiz-Apps. Ich kam von einem auf Textdateien basierenden System (Markdown, um genau zu sein) und für mich war das etwas neues. Mein Arbeiten hat das verändert. Ich bin kein eloquenter Telefontipper, nutze aber Workflowy, um spontane Ideen festzuhalten. Und ich kann meine Notizen in Ad-Hoc-Besprechungen auf dem Telefon konsultieren. Oder die Einkaufsliste im Supermarkt.
Workflowy schreibt natürlich nicht vor, wie die Notizen selbst zu organisieren sind. Ich hatte zunächst ein vereinfachtes, an Getting Things Done angelehntes System, mit nie bereinigtem Eingangsbereich, überquillenden Aufgabenlisten, Ressourcen und so weiter. Ich stieß auf Frank Degenaar mit seinem Zeitstrahlsystem. Er ist ein Workflowy-Veteran und arbeitet, wenn ich das richtig verstehe, auch bei Worklowy, wohl im Support. Ähnlich wie Workflowy selbst glänzt der Zeitstrahl mit konzeptioneller Schlichtheit. Und ich bin immer für einfache Sachen zu haben.
Frank Degenaar erklärt seinen Ansatz am besten in einem YouTube-Video. Er hat auch ein Buch dazu geschrieben. Aber, wenn ich das sagen darf, er ist nicht der größte Literat, das Buch ist überteuert und illustriert beeindruckend meinen Punkt oben, dass Workflowy nicht für Prosa-Texte ist – das Leseerlebnis ist grausam. Nicht zuletzt schreibt er selbst eingangs in dem Buch, dass er nicht verstehe, wozu das Buch nötig sei, wo doch sein System so einfach und offensichtlich ist. Und so ist es auch. Man braucht das Buch nicht.
Herr Degenaar sortiert alle Aufgaben, Projekte, Termine und Ereignisse auf einem Zeitstrahl ein. Dieser besteht aus den Kalendertagen. Es gibt also keinen Eingang, keine Aufgabenlisten, keine Projektlisten. Nur die fortschreitende Zeit, die Tage. (Bei mir gibt es, zugegeben, als Ausnahme neben dem Zeitstrahl noch einen kleinen Bereich für Ressourcen wie Linksammlungen und Tagebücher.) Termine wie zum Beispiel Besprechungen haben auf einem Zeitstrahl naturgemäß einen Platz. Aufgaben und Projekte schiebe ich auf den Tag, an dem ich an ihnen arbeiten oder zumindest an sie erinnert werden möchte. Zu Beginn jedes Tages sortiere ich die vorgemerkten Termine und Aufgaben in den aktuellen Tag ein. Dieser besteht aus den Stunden, die der Tag gibt. Ich halte mich bei der Abarbeitung selten genau an diese Uhrzeiten, aber es gibt dem Tag doch eine grobe zeitliche Struktur und ich werde nicht von einer großen, langen Aufgabenliste erschlagen. Ich kann auch entscheiden, dass eine Aufgabe nicht in den Tag passt und sie auf später verschieben. Gleichsam schiebe ich am Ende des Tages alles nicht erledigte auf einen späteren Tag. In Workflowy geht dieses Einsortieren und Herumschieben fantastisch gut im Wortsinne, nämlich mit der Seitenleiste und per Drag’n’Drop.
Die Einträge, die auf dem Zeitstrahl leben, können beliebig groß und komplex sein. Wenn ich ein Projekt herumschiebe, ist das wirklich das komplette Projekt und nicht nur eine Referenz. Das können Seiten an Aufgaben, Gedanken und Notizen sein, Bilder, Anhänge, vielleicht auch vorhergegangene, zum Projekt gehörende Besprechungen. In Workflowy geht das problemlos von der Hand.
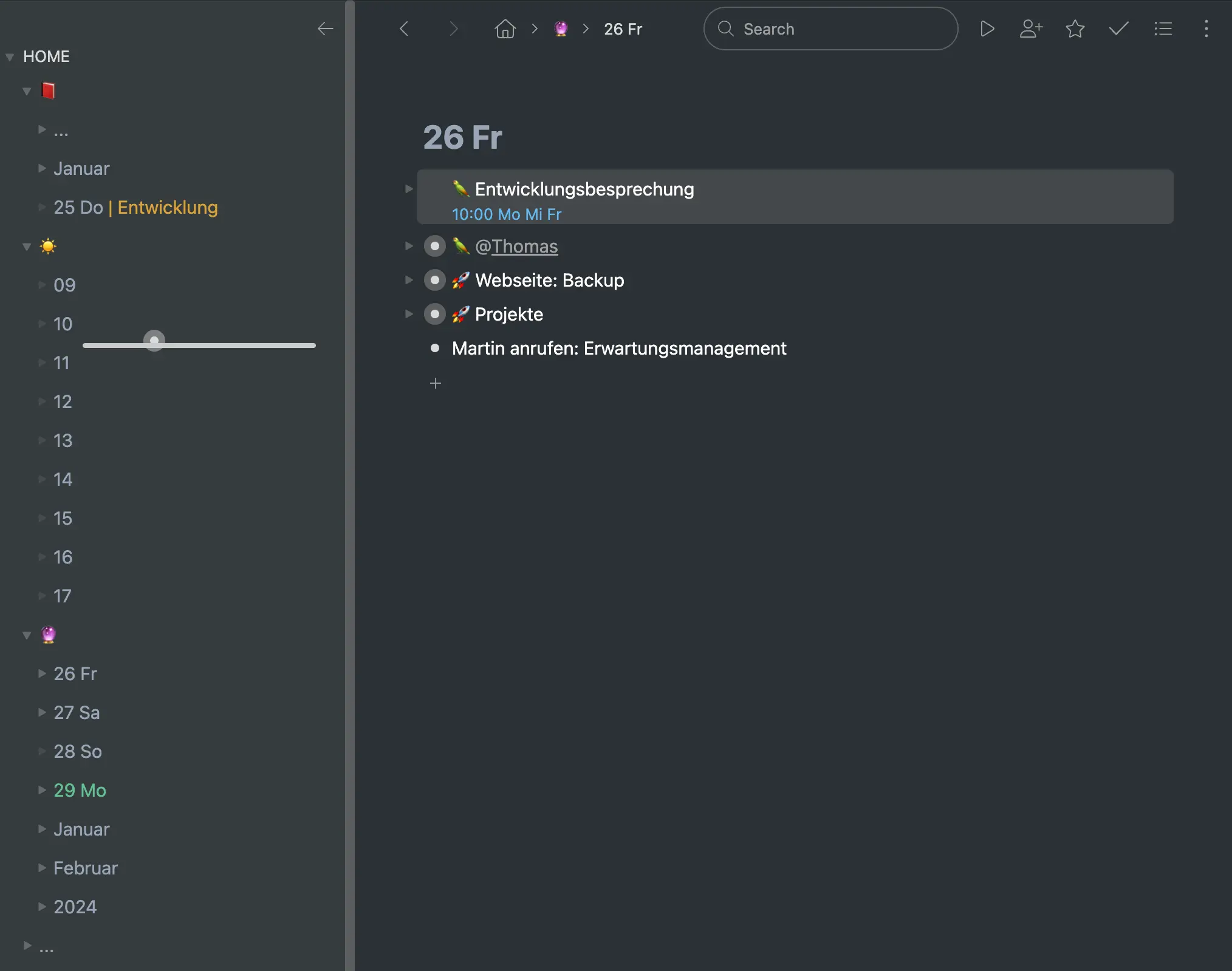
Im Bild beginne ich einen Freitag im Januar. Mein echter Tag geht natürlich nicht nur von neun bis fünf. Und die Aufgabenliste ist real viel länger.
Damit sind die wesentlichen Prozesse im Workflowy Zeitstrahl erklärt. Darüber hinaus gibt es eine Reihe an Details und kleineren Vorgängen. Wiederkehrende Termine – oft wöchentliche Besprechugnen – kopiere ich nach dem Verstreichen auf den nächsten Termin, per Hand. Termine selbst sind an dem blauen Subtext mit der Uhrzeit oder dem Datum zu erkennen. Die Notizen von Besprechungen, gefällte Entscheidungen und Ähnliches kopiere oder verschiebe ich in den vergangenen Tag in der Zeitleiste. (Das ist etwas, was es bei Frank Degenaar nicht gibt – er schlägt vor, diese Dinge zu löschen.) Ich benutze Emoticons und Farben für verschiedene Funktionen. Um ein paar Beispiele zu nennen: Projekte benutzen eine Rakete (🚀), Besprechungen oder Gespräche einen Papagei (🦜), meine Vergangenheit ist ein Buch (📕), der aktuelle Tag eine Sonne (☀️) und die Zukunft eine Glaskugel (🔮). Um Übersicht zu wahren, werden die Tage in Monate hineinsortiert, diese in Jahre, diese unter ”…” zusammengefaltet.
Es ist essentiell, ähnliche Aufgaben oder Punkte zu gruppieren, zusammenzufassen, damit die Listen nicht endlos werden. In Workflowy mit den verschachtelten, faltbaren Listen ist das natürlich ein Leichtes. Bei mir gibt es Gruppen – also Punkte mit Unterlisten – für 🚀 Projekte oder für 🦜 @Thomas (alles, was ich mit Thomas besprechen will), um ein paar Beispiele zu nennen. Die relevante, ja kritische Kehrseite des Zusammenfassens ist, dass man damit wichtige Aufgaben leichter übersehen kann. Vielleicht ist unter 🚀 Projekte etwas, das definitiv an diesem Tag gemacht werden muss – aber da ich die Projekte nicht ausklappe und im Detail besehe, vergesse ich es. Ich schreibe daher manchmal wichtige Unterpunkte mit in die Überschrift hinein – zum Beispiel 🚀 Projekte: Neue Webseite – aber auch das stößt an seine Grenzen.
Wie schon erwähnt kennt der Zeitstrahl keinen dedizierten Eingangsbereich für Dinge – Aufgaben, Ideen, Rückrufe. Stattdessen schreibe ich das, was kommt, dort auf, wo ich gerade bin. Das ist normalerweise die aktuelle Stunde im aktuellen Tag. Spätestens am Ende des Tages (oft aber auch früher, denn ich schaue ja stetig auf den aktuellen Tag) sortiere ich die Notiz weg, als Aufgabe oder Denkanstoß in einen der kommenden Tage.
Was macht nun diesen Zeitstrahlansatz besser als andere Systeme, für mich? Zweifelsfrei ist er konzeptionell einfacher. Es sind auch keine wöchentlichen Reviews und Planungen nötig (die schon bald nicht mehr stattfinden), stattdessen wird täglich und oft mehrmals täglich etwas umentschieden, herumgeschoben. Herr Degenaar nennt das Mikroentscheidungen. Die fortlaufend getroffen werden. In Summe fließt immer noch signifikant Zeit in die Planung der Aufgaben und die Pflege des Systems. Aber sie findet statt, zumindest bei mir. Das System funktioniert. Das liegt wohl auch daran, dass man den neuen Tag erst gut beginnen und also mit Aufgaben und Terminen befüllen kann, wenn man den alten durchgegangen ist und wegsortiert hat.
Das stetige Herumschieben – meist in die Zukunft – von Aufgaben und Projekten ist auf den ersten Blick vielleicht etwas unbefriedigend, entspricht aber meiner Lebensrealität. Der große Vorteil gegenüber anderen Ansätzen ist, dass ich die Aufgaben tatsächlich verlässlich sehe und mir vergegenwärtige. Sie verschwinden nicht unter dem unteren Bildschirmrand in einer langen Todoliste, die ich fast sicher schon nach ein paar Wochen nicht mehr im Detail beschaue. Nicht zuletzt: Wenn man eine kleine Aufgabe das fünfte Mal von einem Tag auf den nächsten verschoben hat, wird es zu dumm und man erledigt sie einfach.
Ebenso kontraintuitiv ist auf den ersten Blick das manuelle Schreiben des Kalenders und das Einpflegen der Termine. Denn dafür gibt es ja die gängigen Kalender-Apps. Praktisch benötigt das Fortschreiben der Kalendertage von Monat zu Monat verschwindend wenig Zeit. Die Termine selbst halte ich tatsächlich doppelt, zum einem im Outlook beziehungsweise iCloud Kalender, zum anderem im Workflowy Zeitstrahl – und das benötigt durchaus extra Zeit, aber immer nur ein paar Sekunden hier und da, nicht am Stück. Für mich ist es das wert, um im Zeitstrahl wirklich alles zu haben und so sicher den Gesamtüberblick zu behalten. Vor allem kann ich so weitere Notizen – die Vorbereitung, ältere Gesprächsprotokolle zum Thema oder gar ganze Projekte – unter dem Termin ablegen. Und nach dem Termin entweder in die Zukunft schieben oder wegarchivieren. Ich habe auch bemerkt, dass das Schreiben der Kalendertage und Einsortieren der Termine etwas seltsam befriedigendes hat. Ich habe so ein besseres Gefühl für die Zeit, habe die Termine präsenter im Kopf und kann mich so besser auf sie einstellen und vorbereiten.
Das ist der Workflowy Zeitstrahl. Kein Notizsystem wird jedoch das Grundproblem lösen, dass es stetig mehr zu tun gibt, als man erledigen kann. Und so ringe ich natürlich auch derzeit mit langen und wachsenden Listen, die in den kommenden Tagen schlummern. Dazu muss ich besser priorisieren und delegieren. Ich weiß das, bekomme es aber nicht hin. Denn Prioritäten setzen bedeutet eigentlich nur, sich einzugestehen, dass man viele Dinge, die man tun möchte oder auch sollte, nie machen wird. Und sie damit gleich zu löschen. Die meisten wirklich wichtigen Dinge – das wissen wir auch – haben die Eigenschaft, dass sie – aus der Aufgabenliste gelöscht oder nicht – stetig wiederkommen. Wieder aus dem Boden der Gedanken und Weltenströme nach oben auf die Tagesordnung steigen.